|
I integrate ecological theory with molecular & computational methods to better understand how host-associated microbiomes are created and maintained. My goal is to use 'omics to better predict beneficial microbial functions (e.g. disease prevention, commensal-pathogen trade-offs, and metabolism) to potentially manipulate microbiome assembly.
Functional Variability in Microbes
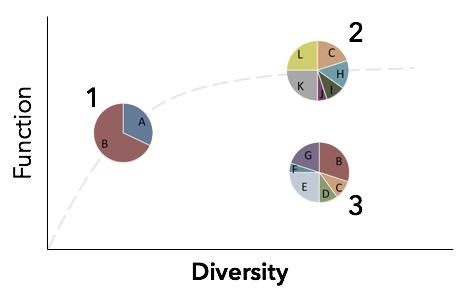
Host-associated microbial communities are usually classified solely using the 16S rRNA gene. As others note, this gene alone is not a reliable indicator of the functional ability of a microbiome partly because genomic diversity is high and adaptation to the local habitat can happen rapidly creating sequence-discrete populations. Thus, 16S data may underestimate functional ability of microbiomes, due to strain-level adaptations to local conditions, or overestimate functional ability, due to functional redundancy across taxa. In my research, I combine taxonomic, genome-resolved metagenomics, and functional analyses to uncover the molecular and metabolic mechanisms of microbe-microbe and host-microbe interactions. Assembly of Host-Associated Microbiomes
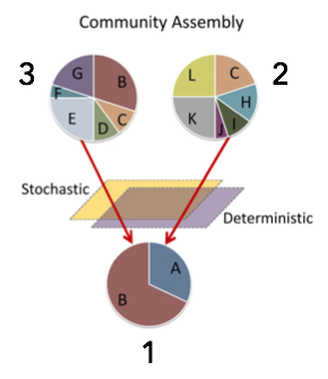
With increasing globalization and environmental disturbance, many organisms may experience variation in the quality of habitat they have available and further the microorganisms they come in contact with. For this reason, I explore the role of both the environment and disease in regulating microbial composition. From environment-to-host, I use molecular and physicochemical techniques to model assembly from the regional pool to local communities to quantify the relative importance of stochastic and deterministic mechanisms. Priority Effects & Probiotics
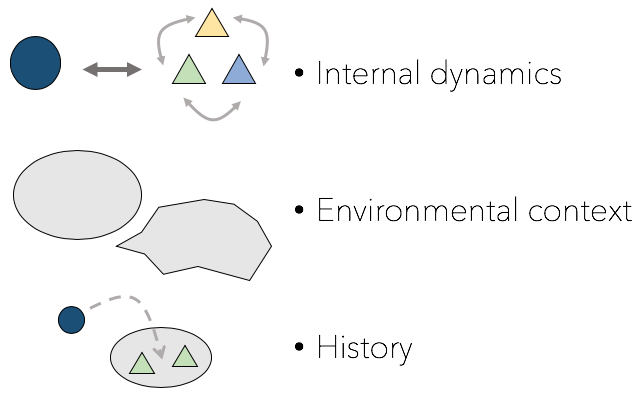
Competition over space and resources is common among microbes. However, the impacts of these interactions are not always predictable based on species identity, niche, and fitness differences, but instead can be historically contingent resulting in priority effects. I use microbial transplant experiments to explore how historically contingent assembly processes can result in a long-term change in microbiome composition creating alternative states of disease prevention. These results have implications not only for which microbes we use in probiotics, but also how they are delivered to hosts. |
(Above images modified from Bannar-Martin et al. 2018)
|